In the first few months of 2025, the CenGen team sequenced the whole genomes of two local malt barley varieties with more to follow – a first for South Africa. A locally conceived project to sequence a South African wheat variety was concluded in 2022, with all the sequencing performed abroad. This revealed the required information to identify the exact DNA letters of a specific wheat disease resistance gene.
The CenGen team and research partners are conducting this barley study with financial assistance from the SA Winter Cereal Industry Agency (SAWCIA), using Oxford Nanopore Technologies (ONT) for sequencing with instrumentation provided by the Department of Science and Technology Innovation through DIPLOMICS (www.diplomics.org.za) (Photo 1).
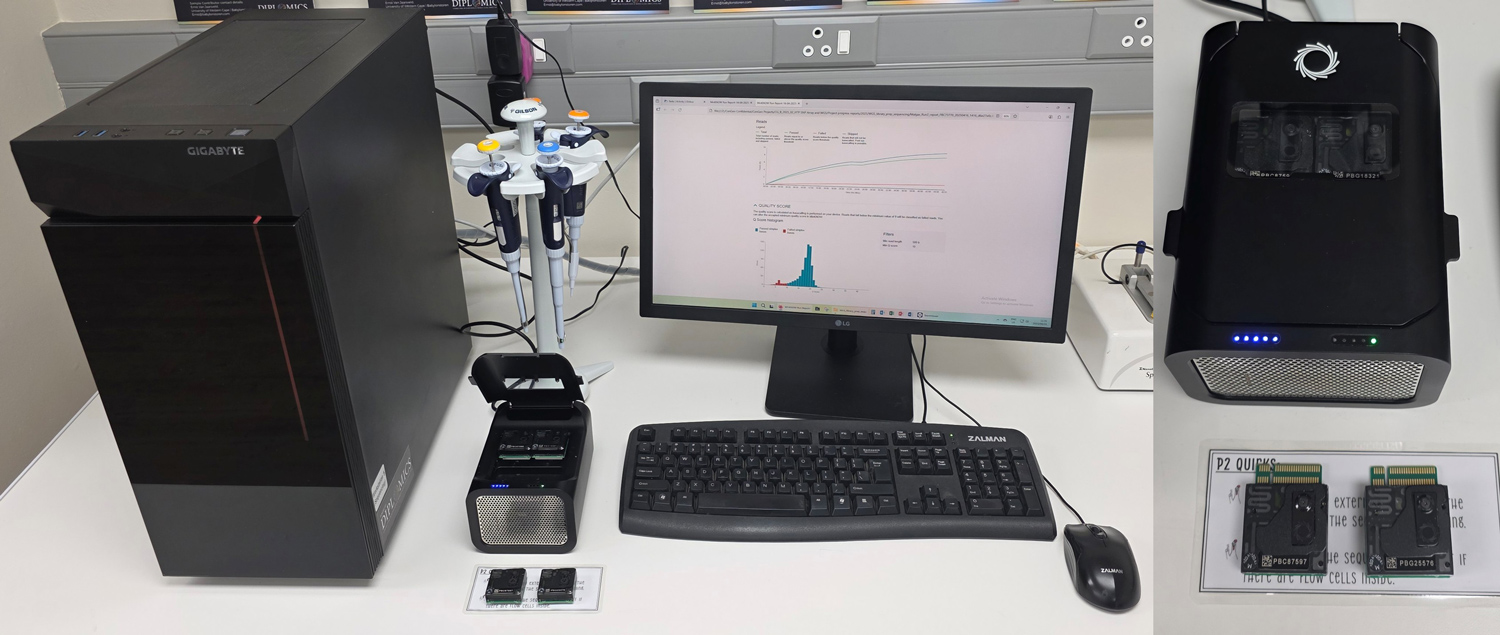

This instrumentation was placed at CenGen in 2024, and all the sequencing work is performed in house. In this study, CenGen aims to determine almost 5 billion DNA letters (base pairs) and their exact order in four important malt barley varieties. Some of these are the founding parents of many of the current commercial varieties.
The first step to understand the whole genome of an important commercial variety is to sequence it, which allows researchers to have the variety laid out in a manner similar to reading a book (Quest 18 (No 2) 24-27, 2022. Pushing the boundaries of SA wheat genetics2). By reading the words organised in a sensible format, we then start to understand the meaning.
There are more than 76 barley genomes (‘books’) that have been sequenced and published by international researchers. Now that the content (order of DNA letters) of two South African barley varieties is known, CenGen can compare it with the genomes of barley varieties from across the world. This will help us to study specific genes that code for important characteristics/traits of the varieties with a focus on malt barley. The whole genome sequence is key to deciphering all this information.
How the genes manifest or are expressed in a variety is, amongst other factors, determined by the allele (specific form of the gene) represented in that variety. The international barley community has put in a huge effort to also determine which of these DNA letters code for which genes. In Figure 1, some of the essential traits, and the known genes coding for them are highlighted. For instance, culm length is an important agronomic trait and South African irrigation and dryland varieties typically have optimal heights of ~80 cm and ~70 cm, respectively. In the current study, preliminary comparisons have indicated that the SDW1.d allele of the SDW1 gene (coding for dwarf, semi-dwarf, and medium-tall plants) is present in Nemesia and Malgas, confirming which allele/form of the gene is required in South African malt barley to have plants within the range that the industry prefers. The research team is now systematically working through all these genes and determining which alleles are present in our barleys.
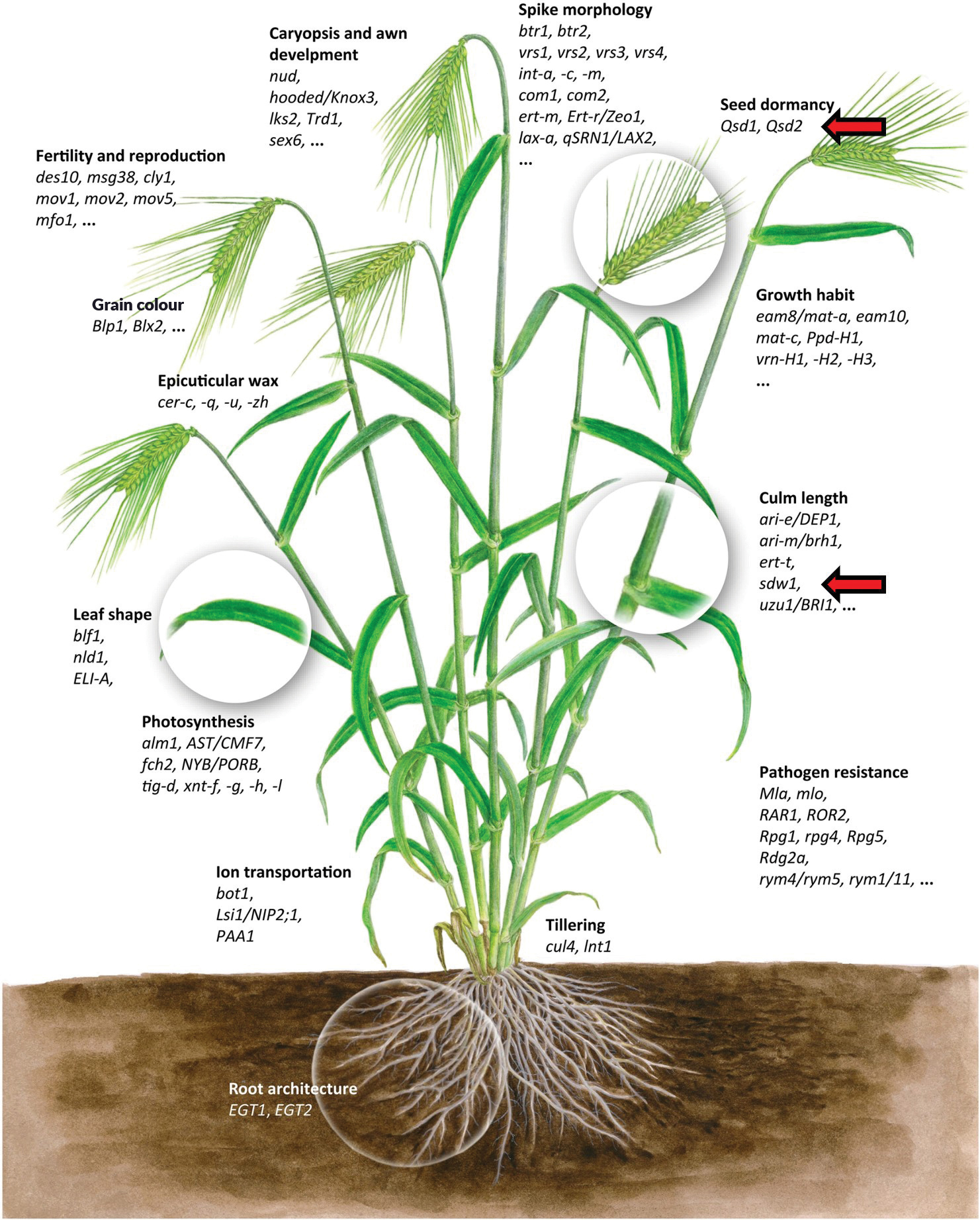
Having the whole genome sequence, and how it compares to those of international varieties, provides insight to our local barley breeders on a level they have never had before. This approach will assist in identifying solutions to current issues faced by the South African barley industry while keeping in mind challenges experienced worldwide. One can then pre-emptively identify solutions and breed selectively to incorporate desirable alleles, to be ready should these problems arise locally.
CenGen also works closely with the South African Barley Breeding Institute (SABBI) and Dr Burkhard Steuernagel, head of the Informatics Group at John Innes Centre (JIC) in the United Kingdom. Wilku Meyer, research scientist: Genomics and Bioinformatics, recently spent a month at the JIC to specifically study these large data sets3. The long-standing collaboration between CenGen and the JIC has benefitted the whole small-grain industry over many years.
References